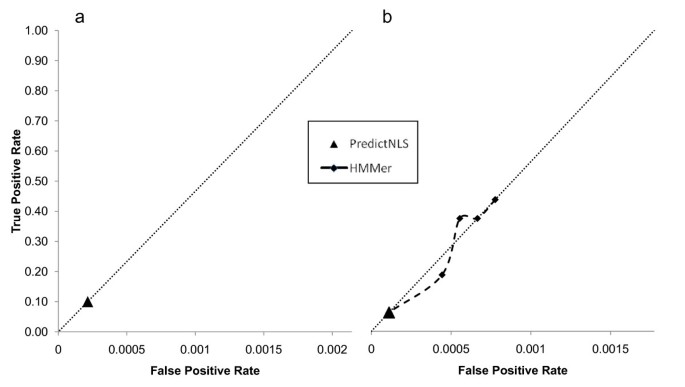
NLStradamus: a simple Hidden Markov Model for nuclear localization signal prediction | BMC Bioinformatics | Full Text

The flow charts of predicting NLS. (a) The flow chart of mining the... | Download Scientific Diagram

Nuclear Localization Signal | In Silico Nuclear Localization Signal Prediction | NLS Mapper| - YouTube

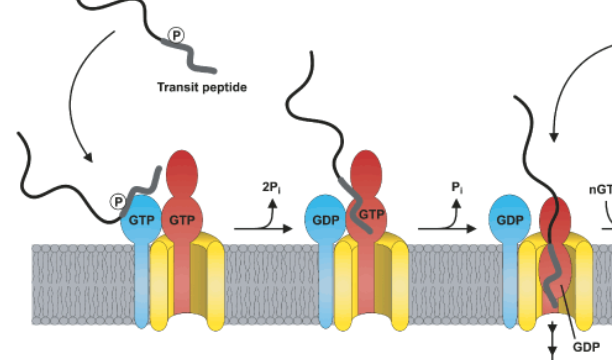
Identification of a nuclear localization signal in the Plasmodium falciparum CTP: phosphocholine cytidylyltransferase enzyme | Scientific Reports

Biology | Free Full-Text | Piggybacking on Classical Import and Other Non-Classical Mechanisms of Nuclear Import Appear Highly Prevalent within the Human Proteome

Figure 2.1 from Nuclear export signals (NESs) in Arabidopsis thaliana : development and experimental validation of a prediction tool | Semantic Scholar

Nuclear Localization Signal | In Silico Nuclear Localization Signal Prediction | NLS Mapper| - YouTube
SeqNLS: Nuclear Localization Signal Prediction Based on Frequent Pattern Mining and Linear Motif Scoring | PLOS ONE

Delivery of eGFP fused with nuclear localization signals (NLS) to cells... | Download Scientific Diagram
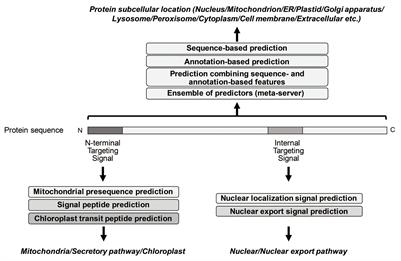
Frontiers | Tools for the Recognition of Sorting Signals and the Prediction of Subcellular Localization of Proteins From Their Amino Acid Sequences

Software tools for simultaneous data visualization and T cell epitopes and disorder prediction in proteins - ScienceDirect

Discovering nuclear targeting signal sequence through protein language learning and multivariate analysis - ScienceDirect
NES and NLS prediction in the bHLH domain of TCF4. (A) Results of NES... | Download Scientific Diagram
SeqNLS: Nuclear Localization Signal Prediction Based on Frequent Pattern Mining and Linear Motif Scoring | PLOS ONE

NLS Mapper, PredictProtein and COMPARTMENT OAS3 nuclear localization... | Download Scientific Diagram

Cells | Free Full-Text | Bioinformatics and Functional Analysis of a New Nuclear Localization Sequence of the Influenza A Virus Nucleoprotein

NLStradamus: a simple Hidden Markov Model for nuclear localization signal prediction | BMC Bioinformatics | Full Text